This research theme aims to discover and characterise natural microbial defence systems with the potential to be used to explore pest/pathogen biology and address bioprotection challenges.
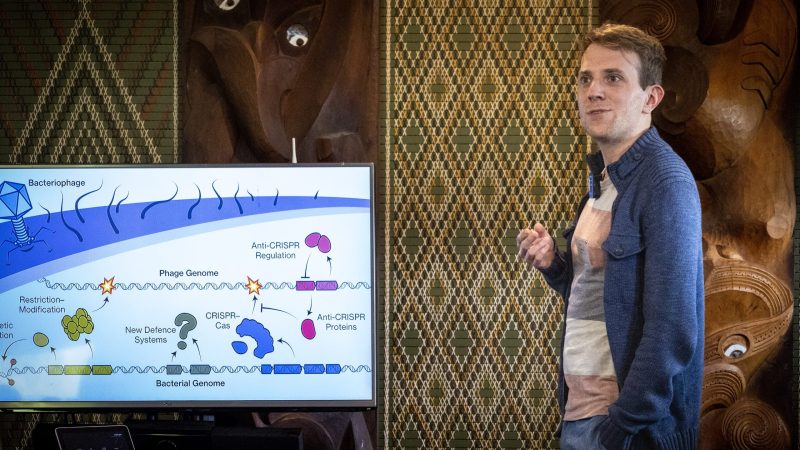
Molecular technologies derived from bacterial defence systems (e.g. restriction enzymes and CRISPR) have a proven track record in yielding powerful technologies. CRISPR-Cas can be programmed to recognise – and can modify – specific sequences, making them adaptable to study pest biology, as diagnostics, and sequence-specific biocontrol agents to eliminate particular bacteria.
Bacteria also have a multitude of other defence systems and many new ones are being discovered. Aotearoa New Zealand microorganisms are likely to harbour novel defence systems that can be applied for innovative research and bioprotection strategies. This work will benefit from sequencing data and insight from the research out of 1.2 Interconnected Properties and other projects where microorganisms are studied.
We are combining bioinformatic and experimental approaches to discover and study defence systems from bacteria. This work will reveal the potential of any of these systems for future development as tools for Bioprotection research and their possible influence on biocontrol approaches that rely on targeted bacterial killing by bacteriophages (viruses that infect bacteria). We will also work with Aroha Mead (Recloaking Papatūanuku team) to explore Māori views and best practice for biodiscovery work and access and benefit sharing.
We have uncovered that some bacteria contain multiple defence systems that can antagonise each other (Birkholz et al., 2022 NAR). We have also further developed bioinformatic approaches for the detection of known phage defence systems in bacterial genomes (Payne et al., 2021 NAR). We have identified new potential defence systems for further testing and are exploring the biotechnological potential of other systems we are studying.
Research Projects
2.3 Harnessing Biodefences
Fighting crop pathogens with viruses
Project Team